BioWire Weekly - 028
Biotech News
Happy Monday Morning, Readers. Let’s be relentless this week!
Okay, let’s address the elephant (or mammoth?) in the room, I’m a bit behind on the newsletters. Sadly, its not because I’m off adventuring or enjoying a much-needed vacation somewhere. I’ve been navigating some personal challenges related to a relationship. Maybe someday I’ll even unpack those experiences here, but for now, I just appreciate your patience while I rediscover my rhythm.
If we’re being honest, BioWire Weekly should probably be BioWire BiWeekly. Jokes aside, I genuinely enjoy every aspect of making this content - the research, writing, and especially adding my personal take. It’s a much needed outlet. Realistically, though, a two-week cadence is probably what’s most sustainable for these roundups in the near term. But hey, If a big sponsorship deal rolls in, I’ll gladly revisit this!
At that, let’s dive in.
The topics this week:
A Woolly Mouse represents a tiny, but mighty, step toward a Mammoth
A Meditation on Making Superbabies
Sparking life in a flask
Can Omega-3 Slow Down Aging? What Epigenetic Clocks Say.
Evo 2 is AI for Genome Engineering at an Unprecedented Scale
In cancer, can the medicine be worse than the disease?
If you enjoy these updates, consider subscribing and becoming a part of our growing community!
A Woolly Mouse represents a tiny, but mighty, step toward a Mammoth
Okay, this news is too exciting not to share, but I am a bit biased here. Why? A little over two years ago, I moved to Texas to join Colossal Biosciences, the biotech startup famously aiming to bring back the extinct mega-fauna, the woolly mammoth.
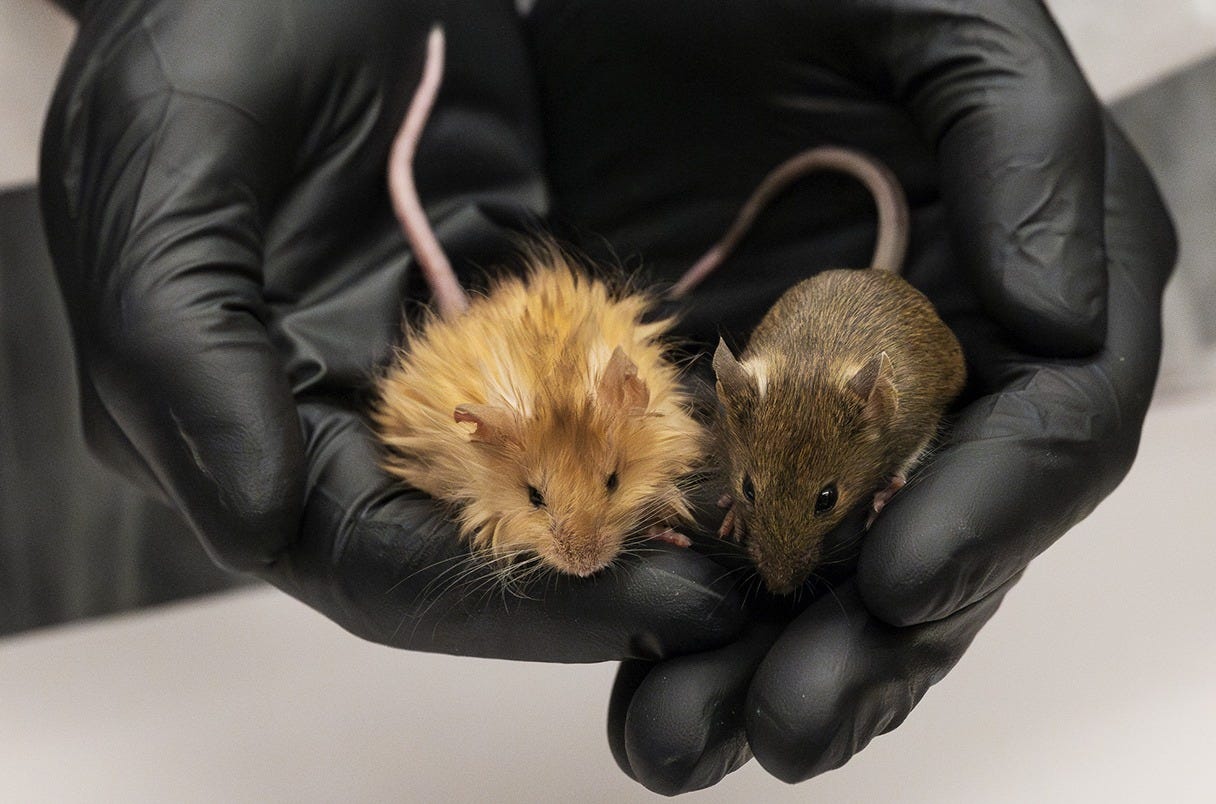
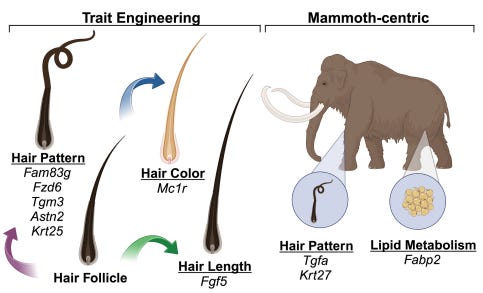
Last week, Colossal had a big announcement: the successful creation of engineered mice carrying woolly mammoth traits, what is now referred to as the woolly mouse! The genes targeted were associated with hair growth and metabolism, producing mice with thick, fluffy coats that genuinely reassemble miniature woolly mammoths. The pictures are absolutely striking - it’s rare for scientific progress in biotechnology to be so completely communicated in one image. Moreover, they’re shockingly cute.
If you can look beyond the adorable mouse though, there are some heavy things to unpack. This feat was achieved with advanced multiplex gene editing technology. These mice are just one demonstration. You can only imagine what is around the corner, particularly for their ability to revive lost traits and species. For those curious, you can dive deeper into the details in Colossal's recent publication (Chen et al., 2025). But fair warning: you might find yourself wishing for your own pet woolly mouse.
A Meditation on Making Superbabies
This week, I came across a provocative and fascinating deep-dive article on LessWrong about the future of genetic enhancement, something the author playfully refers to as the creation of "Superbabies." The author Gene Smith (clever pseudonym) details the current state and future possibilities of embryo selection and gene editing technologies. Honestly, the article feels a bit like science fiction—Gattaca is probably what comes to mind. But hey, I like science fiction!
How does in vitro fertilization (IVF) work and what is embryo selection? We’ve had several articles outlining the IVF process, including the process and emerging technologies that are aimed at helping couples with fertility issues (read more here - link 1, link2). As a quick reminder, IVF involves the extraction of oocytes (egg cells) from a woman, their fertilization with sperm outside of the body, and their maturation to the blastocyst stage for implantation. However, when this is performed, it’s not just 1 embryo. Typically, 10-15 embryos are matured at a time, since often the first round of IVF is not successful. These additional embryos are frozen and kept for future IVF attempts.
Could there be a benefit of choosing specific embryos for implantation based on their genetic makeup? Right now, embryo selection companies like Heliospect claim that they can already offer parents a slight edge by helping them choose embryos based on traits like intelligence or disease resistance, potentially adding around 3-7 IQ points or a few healthy years to a child's life. A simulation of these enhancements through embryo selection based on available embryos seems to indicate that improvement could be made with a greater selection, but overall, this levels off fairly quickly. The authors claim that to make real leaps, they would need to directly edit genes.
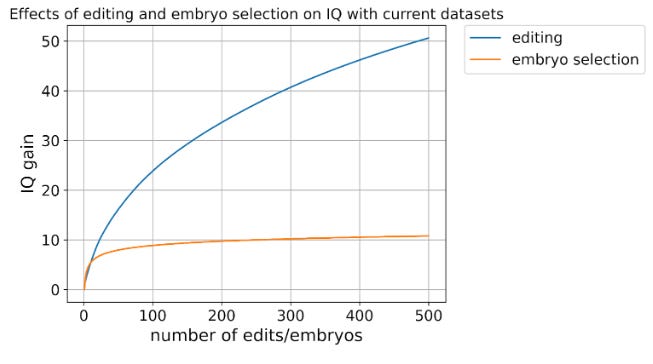
Gene editing, especially with powerful new methods like CRISPR, could theoretically boost IQ by up to 50 points or extend lifespans by decades by carefully tweaking just a few key genetic sites. But there's an even bigger breakthrough looming: researcher Sergiy Velychko recently developed something called "Super-SOX," a technique that dramatically improves our ability to reprogram cells into embryonic stem cells. This isn't a minor improvement—it boosted the survival rates of edited mouse embryos by an incredible 800%, suggesting that extensive, precise genetic edits might soon become feasible at scale.
Of course, as exciting as this sounds, huge questions remain—not only about the safety and long-term effects but also the ethical and social implications. Human germline editing (making heritable genetic changes) is currently off-limits in many countries, including the U.S., due to ethical and regulatory concerns. And that's probably for good reason. Tweaking our genetic code to enhance human traits is likely to open a Pandora's box of unforeseen consequences.
Despite the many hurdles and ethical questions, it's hard not to get a little excited (or at least intrigued) by the possibilities here.
Sparking Life in a Flask
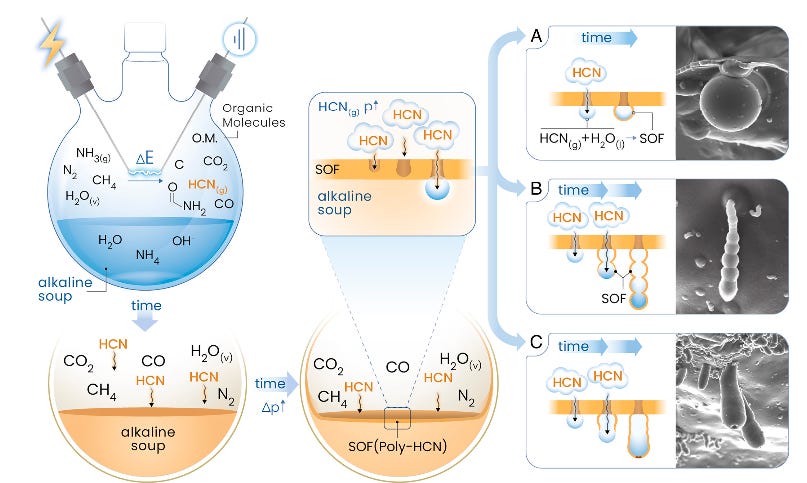
The boundary between chemistry and biology has always been shrouded in mystery—where, when, and how did life emerge from lifeless matter? A new study published in PNAS offers a compelling glimpse into this transition, demonstrating that protocells and key prebiotic molecules could have formed simultaneously under conditions plausible on the early Earth (Jenewein et al., 2025).
Using a modified Miller-Urey experiment, researchers recreated an ancient reducing atmosphere - rich in methane (CH₄), ammonia (NH₃), nitrogen (N₂), and water vapor - and subjected it to electrical discharges, mimicking lightning strikes on the primordial Earth. The outcome was striking: not only did the experiment generate a diverse set of prebiotic molecules, but it also led to the spontaneous formation of protocells—hollow, vesicle-like structures that could have acted as microreactors for early biochemical evolution. These protocells emerged within a silica-induced organic film, demonstrating that mineral surfaces may have played a crucial role in fostering the origins of life.
A key aspect of this study is its challenge to conventional prebiotic models. Traditionally, scientists have treated the synthesis of life’s molecular building blocks (e.g., amino acids, nucleotides) and cellular compartmentalization as distinct events, separated in time and space. Here, however, both processes occurred simultaneously, suggesting that life’s emergence may have been more spontaneous and inevitable than previously thought. The protocells formed via a bubble-driven mechanism, in which polymerized hydrogen cyanide (HCN) self-assembled into hollow biomorphic structures at the water-gas interface. These compartments could have concentrated prebiotic molecules, setting the stage for self-replicating chemical systems.
The implications extend far beyond Earth. If such conditions can generate protocells and prebiotic chemistry on any planet with water and a similar atmospheric composition, then the emergence of life may not be a rare cosmic event, but a fundamental chemical process. This work also raises new challenges for astrobiology and life detection efforts, as the discovery of hollow, biomorphic structures on Mars or in ancient terrestrial rocks may not necessarily be evidence of past life, but rather a byproduct of abiotic chemical processes.
Can Omega-3 Slow Down Aging? What Epigenetic Clocks Say.
The pursuit of longevity often revolves around high-tech interventions, but what if a simple nutritional supplement could make a measurable impact? A new study from the DO-HEALTH trial, published in Nature Aging, suggests that omega-3 supplementation may decelerate biological aging, as measured by DNA methylation (DNAm) clocks (Bischoff-Ferrari, et al., 2025).
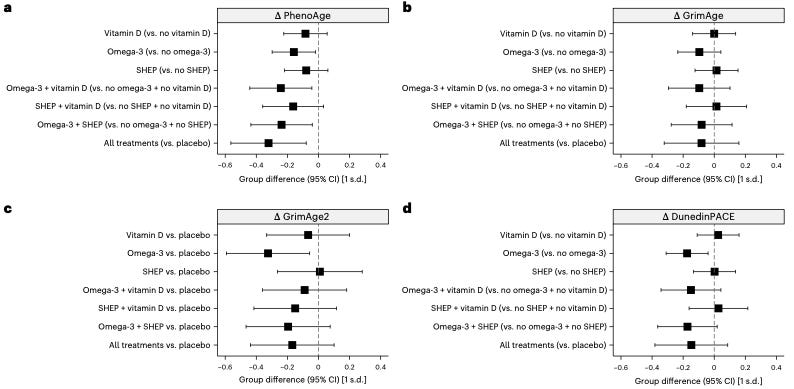
Epigenetic clocks, such as GrimAge and PhenoAge, are powerful biomarkers that estimate biological age by analyzing DNA methylation patterns. In this randomized controlled trial, 777 older adults received daily doses of omega-3 (1g), vitamin D (2,000 IU), and/or a structured home exercise program over three years. The results? Omega-3 supplementation alone significantly slowed aging markers on three separate epigenetic clocks—PhenoAge, GrimAge2, and DunedinPACE. More interestingly, when combined with vitamin D and exercise, the benefits on PhenoAge were even more pronounced. However, vitamin D and exercise alone did not show consistent effects across the clocks.
While the effect sizes were modest—equating to about 2.9 to 3.8 months of biological age deceleration over three years—the findings reinforce the idea that small, sustained interventions can accumulate meaningful longevity benefits over time. Omega-3’s effects appeared strongest in individuals with lower baseline levels of DHA and EPA, suggesting that personalized approaches to supplementation could optimize results.
This study adds to a growing body of evidence that supports omega-3 as a viable, low-risk geroprotective intervention. However, whether this translates into tangible increases in lifespan or healthspan remains an open question. Future research with longer follow-up periods will be crucial in determining whether these epigenetic shifts correspond to real-world longevity benefits.
Evo 2 is AI for Genome Engineering at an Unprecedented Scale
The biological world is increasingly being shaped by AI, and the release of Evo 2 represents a milestone in genomic modeling (Brixi et al., 2025). Developed by researchers at the Arc Institute and Stanford University, Evo 2 is a foundational AI model trained on 9.3 trillion DNA base pairs, spanning genomes from across all domains of life. Unlike previous models, which were largely limited to analyzing protein sequences or short genomic segments, Evo 2 has been designed with an unprecedented 1 million base pair context window, allowing it to model entire genomes with remarkable precision.
Evo 2 operates as both a predictive and generative model. It can assess the functional impact of genetic variations, including noncoding mutations, pathogenic variants, and BRCA1 risk factors—without any task-specific fine-tuning. This zero-shot capability makes it a powerful tool for predicting disease-causing mutations and refining variant classification. More impressively, Evo 2 is capable of designing entire genomes, generating mitochondrial, prokaryotic, and even eukaryotic sequences with coherence and biological realism. Early tests suggest that Evo 2 outperforms previous models in simulating complex biological structures, such as chromatin accessibility, which governs gene expression.
Beyond its technical capabilities, what sets Evo 2 apart is its commitment to openness. Unlike many AI models in biology that remain proprietary, Evo 2 has been fully open-sourced, including model weights, training code, and the vast OpenGenome2 dataset. This transparency is a rarity in AI-driven genomics and has the potential to accelerate research in synthetic biology, precision medicine, and evolutionary studies. However, with such power also comes scrutiny - particularly regarding safety. The developers have proactively excluded viral sequences that infect humans from Evo 2’s training set to prevent misuse, though concerns about potential risks remain.
Ultimately, Evo 2 represents a significant leap in biological foundation models. By merging large-scale AI with the complexity of life's genetic code, it opens the door to a future where genome design could be as programmable as software engineering. Whether this leads to breakthroughs in synthetic biology or raises new bioethical challenges, one thing is certain: AI-driven biology is accelerating faster than ever before.
In cancer, can the medicine be worse than the disease?
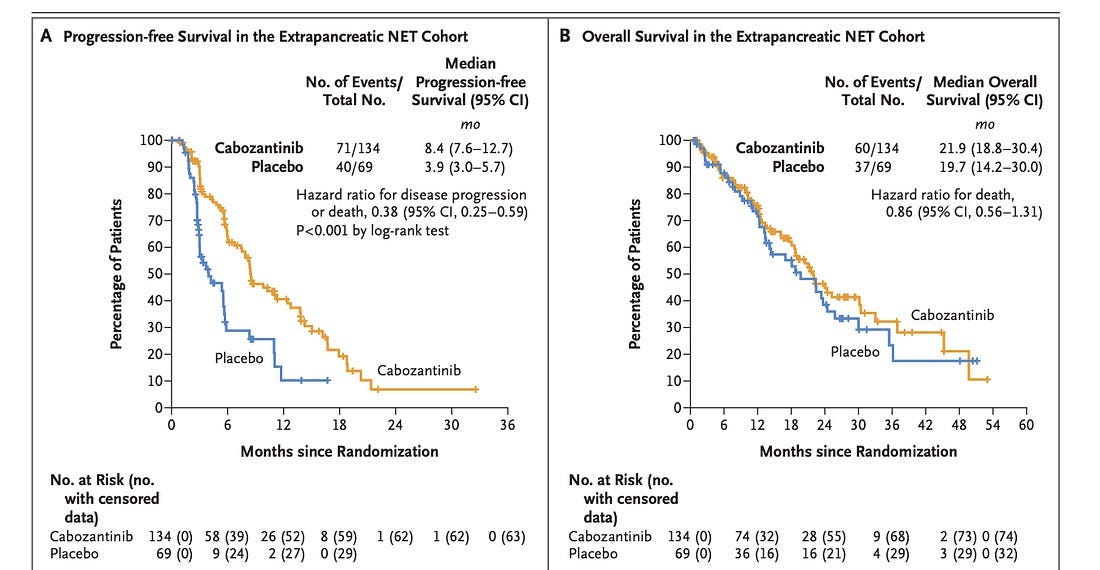
A newly published New England Journal of Medicine study has reignited debate over oncology trial design and the real-world utility of targeted therapies. The CABINET trial, funded by the National Cancer Institute (NCI), investigated cabozantinib, a multi-kinase inhibitor, in patients with advanced neuroendocrine tumors (NETs) (Chan, et al., 2024). The randomized, placebo-controlled phase 3 trial enrolled 298 patients with progressive pancreatic or extra-pancreatic NETs, assigning them 2:1 to either cabozantinib (60 mg daily) or placebo. The results were statistically significant: cabozantinib extended progression-free survival (PFS) from 3.9 months to 8.4 months in extra-pancreatic NETs and from 4.4 months to 13.8 months in pancreatic NETs. However, despite delaying tumor progression, the drug failed to improve overall survival (OS), raising critical questions about its true clinical impact. Furthermore, toxicity was substantial, with 62-65% of patients experiencing grade 3 or higher adverse events, including hypertension, fatigue, and diarrhea.
At face value, the trial suggests cabozantinib could be a new treatment option for progressive NETs. But its design and methodology have drawn sharp criticism. Oncologist Vinay Prasad and others argue that the choice of a placebo control rather than the best available standard of care - such as Lu-177 dotatate or everolimus - is ethically questionable. Patients in clinical practice would not be left untreated, making the placebo comparison misleading. Another red flag is that midway through the trial, the study switched from centralized radiographic review to local assessments. This may have biased progression assessments, making cabozantinib appear more effective than it truly was. Additionally, the 2:1 randomization ratio, rather than the standard 1:1, reduces statistical power while making the trial more attractive for recruitment.
The biggest problem, however, lies in the reliance on progression-free survival as a primary endpoint. PFS is an imperfect surrogate for patient benefit, often failing to translate into longer life expectancy or improved quality of life. Many argue that oncology trials need to prioritize overall survival and patient-reported outcomes, rather than merely demonstrating a delay in tumor growth. If cabozantinib does not help patients live longer, and if it adds substantial toxicity, what are we really gaining? This case underscores a broader issue in drug approval: whether pharmaceutical companies are incentivized to push costly, marginally effective treatments based on metrics that do not always align with meaningful patient outcomes.
Ultimately, cabozantinib’s role in neuroendocrine tumors remains uncertain. While it may slow disease progression, its failure to improve survival, its substantial toxicity, and the questionable trial design leave its true clinical value up for debate. As targeted cancer therapies continue to emerge, the challenge remains to ensure that drugs aren’t just approved based on technical success in trials—but that they deliver real, tangible benefits for patients.
Support:
These newsletters take a significant amount of effort to put together and are totally for the benefit of the reader. If you find these explorations valuable, there are multiple ways to show your support:
Engage: Like or comment on posts to join the conversation.
Subscribe: Never miss an update by subscribing to the Substack.
Share: Help spread the word by sharing posts with friends directly or on social media.
References:
Bischoff-Ferrari, H.A., Gängler, S., Wieczorek, M., Belsky, D.W., Ryan, J., Kressig, R.W., Stähelin, H.B., Theiler, R., Dawson-Hughes, B., Rizzoli, R. and Vellas, B., 2025. Individual and additive effects of vitamin D, omega-3 and exercise on DNA methylation clocks of biological aging in older adults from the DO-HEALTH trial. Nature Aging, pp.1-10.
Brixi, G., Durrant, M.G., Ku, J., Poli, M., Brockman, G., Chang, D., Gonzalez, G.A., King, S.H., Li, D.B., Merchant, A.T. and Naghipourfar, M., 2025. Genome modeling and design across all domains of life with Evo 2. bioRxiv, pp.2025-02.
Chan, J.A., Geyer, S., Zemla, T., Knopp, M.V., Behr, S., Pulsipher, S., Ou, F.S., Dueck, A.C., Acoba, J., Shergill, A. and Wolin, E.M., 2024. Phase 3 Trial of Cabozantinib in Advanced Neuroendocrine Tumors. The New England journal of medicine, 392(7), p.653.
Chen, R., Srirattana, K., Coquelin, M.L., Vilar Sampaio, R., Wilson, R., Ganji, R., Weston, J., Ledesma, A., Beebe, J., Sullivan, J. and Qin, Y., 2025. Multiplex-edited mice recapitulate woolly mammoth hair phenotypes. bioRxiv, pp.2025-03.
Jenewein, C., Maíz-Sicilia, A., Rull, F., González-Souto, L. and García-Ruiz, J.M., 2025. Concomitant formation of protocells and prebiotic compounds under a plausible early Earth atmosphere. Proceedings of the National Academy of Sciences, 122(2), p.e2413816122.
https://www.lesswrong.com/posts/DfrSZaf3JC8vJdbZL/how-to-make-superbabies?utm_source=substack&utm_medium=email






Wow, bumper crop of stories and info here David. Thanks.
This is wild: "This isn't a minor improvement—it boosted the survival rates of edited mouse embryos by an incredible 800%, suggesting that extensive, precise genetic edits might soon become feasible at scale."
Also, huge thanks for the PNAS paper. That one is going straight to some of my students who have just been learning about amino acids and the original Miller-Urey experiment.
Would you rather fight 100 mouse sized Wolly Mammoths, or 1 Wolly mammoth sized mouse?